FastAmp® Plant Direct PCR Kit is suitable for amplification of DNA directly from plant samples without purifying DNA
FastAmp® Plant Direct PCR Kit
Price range: $155.00 through $510.00
Description
Intact Genomics FastAmp® Plant Direct PCR kit is an easy and robust method to amplify DNA fragments from plant tissues with high specificity and high sensitivity without the need of complicated DNA purification steps. The advanced formulation of this kit also allows fast PCR cycling conditions without compromising PCR sensitivity, specificity and yield. The FastAmp® PCR kit is an ideal and powerful tool for high-throughput genotyping, DNA amplification, and plant genome analysis.

FastAmp® Plant Direct PCR Kit is suitable for amplification of DNA directly from plant samples without purifying DNA. This kit is based on specially engineered Taq DNA polymerase, proprietary buffer system, dNTP, MgCl2 ,PCR facilitators and dye mix which makes it extremely robust and tolerant of plant PCR inhibitors such as complex polysaccharides, polyphenols and others. This kit has been tested with leaves and seeds from a wide variety of plant species. This PCR Kit includes a complete set of optimized reagents and detailed protocols making it an ideal choice for amplification of plant DNA without DNA purification.
Highlights
- Direct PCR- no need to purify DNA
- Specially engineered Taq DNA polymerase with highest sensitivity and specificity
- Extremely short PCR protocol times
- Master mix format with premixed gel loading dye to reduce cross-contamination and sample handling
errors - Optimized for both low and high GC templates
Applications
- Genotyping
- Transgene detection
- Knockout analysis
- Sequencing
Product Includes
- FastAmp® Plant Direct PCR master mix (2x)
- FastAmp® Plant Direct PCR/Genotyping Solution
Storage Temperature
–20ºC
Quality Control Assays
This PCR Kit has been tested with tissue from a wide variety of plant species, some of the results are included here.
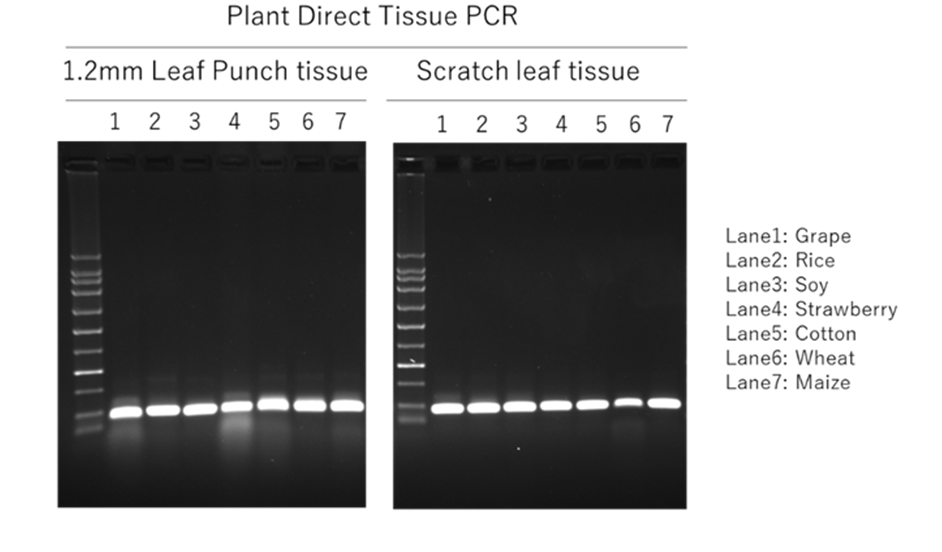
Direct PCR analysis from various 1.2mm punched leaf tissue and scratch tissue using FastAmp® Plant Direct PCR kit
4612 4615
Additional information
| Reactions | 250 Reactions, 1250 Reactions |
|---|
General Guidelines before start
A. Sample handling
1.2mm punch leaf tissue should be placed directly into 8µl FastAmp® Plant Direct PCR/Genotyping Solution in PCR tube.
A leaf punch can be obtained by placing the leaf puncher (i.e. Uni-Core punch by GE Healthcare, Product # WB100028, or any other commercial sources) in perpendicular position over the expanded leaf and rotating it. The tip of puncher needs to placed inside the PCR tube and expunge to drop the sample. It is critical that plant tissue materials/leaf punch is completely inside the PCR solution in tube. Occasionally, the leaf punch may stick to the side of PCR tube and therefore does not reach the bottom of tube. If this occurs, use a 10 µl pipet tip to manually drive the leaf punch to the bottom of PCR tube. This can be done right before capping the PCR tubes and starting the thermocycling program. For reusing the puncher, it is very important to clean the cutting edge properly with 70% ethanol to prevent cross-contamination between samples.
B. PCR conditions
B-1 Denaturation:
An initial denaturation of 8 minutes at 95°C is sufficient for most amplicons. Longer denaturation times can be used (up to 10 minutes) for difficult templates. During thermocycling, the denaturation step should be kept to a minimum. Typically, a 20–30 second denaturation at 95°C is recommended for most templates.
B-2 Annealing:
Optimize the annealing temperatures for the target gene specific amplification by keeping annealing temperature at least 5 ºC below Tm values. Typically, use a 10–30 second annealing step. A temperature gradient can also be used to optimize the annealing temperature for each primer pair.
B-3 Extension:
The recommended extension temperature is 72°C. Extension times are generally 1 minute per kb for complex, genomic samples, but can be reduced to 30 seconds per kb for simple templates. When amplifying products >2 kb, it is often helpful to increase the extension time.
A final extension of 5 minutes at 72°C is recommended.
B-4 Cycle number:
Generally, 35–40 cycles yield sufficient product.
B-5 Primers:
Forward and reverse primers are generally used at the final concentration of 0.1-0.6 µM each. If the primer
concentration is too high, the specificity of priming may be reduced, resulting in non-specific products.
B-6 PCR product:
The PCR products generated using Taq DNA Polymerase have dA ends. If cloning is the next step, then T/A-cloning is preferred.
Protocol
- Thaw 2x master mix, Plant Direct PCR/Genotyping Solution, primers and mix thoroughly and spin down before use.
- Place scratch leaf tissue or punch leaf tissue into 8µl FastAmp® Direct PCR/Genotyping Solution and mix thoroughly
- Prepare a reaction mix according to the following table:

- Mix the reaction mixture thoroughly.
- Program the thermal cycler according to the manufacturer’s instructions. A typical PCR cycling program is outlined in the following table.

- Place the PCR tubes in the thermal cycler and start the cycling program.
- Run 10.0 µl of PCR products in 1% agarose gel (120 volts for 45 min).
Troubleshooting